Research - (2022) Volume 10, Issue 8
Identifying Potential Inhibitors of the Human A-amylase Enzyme via Molecular Docking and Virtual Screening
Nursel Eski1, Cengiz Z Altuntas2, Harun Yilmaz3 and Senol Dane4*
*Correspondence: Senol Dane, Department of Physiology, Faculty of Basic Medical Sciences, College of Health Sciences, Nile University of Nigeria, Abuja, USA, Email:
Abstract
Background: The alpha amylase enzyme plays a critical role in starch digestion to glucose, affecting the blood glucose levels present in one’s bloodstream. Objective: Inhibition of this enzyme can, potentially, modify its catalytic activity and treat a patient with diabetes mellitus. By preventing the catalysis of the a-amylase enzyme, the rate at which carbs break down to sugar and enter the bloodstream can be reduced, decreasing the glucose in one’s bloodstream. This can be done by performing molecular docking to test the binding affinity between phytochemicals and the active site of the protein. With the right dosage and concentration, the phytochemicals with the lowest, or best, binding affinity can be considered to later produce a promising drug that goes off as oral treatment that would be taken with one’s diet to slow the reaction rate. As this study is nothing new and has been experimented on before, the goal of this project is to compare the results to those of the experimentations. Methods: By finding the binding affinity of phytochemicals with the enzyme’s active site, it can be determined which of them are favorable. These phytochemicals can be retrieved from plants that have already been scientifically proven to have effects on the human alpha-amylase enzyme. Once docking of the chemical constituents with the protein’s active site is complete, toxicity checking of the phytochemicals can be done through “mcule”. Whether or not the ran phytochemicals contain inhibiting effects can be determined by creating a threshold which is calculated by first running already- known inhibitors of this enzyme, finding their mean binding affinities, and calculating the standard deviation. Results: About 64 out of the 109 phytochemicals ran met the calculated threshold, the phytochemical Taraxasterol, from the plant Taraxacum, yielding the best binding affinity of –12.36 kcal/mol, although the toxicity checker pointed out potentially toxic parts of its structure. Oleanolic acid is a close and apparent nontoxic second, however, with a binding affinity of -11.72 kcal/mol. Conclusion: Surely, most of the phytochemicals ran have remarkable binding affinities with the active site of the alpha- amylase enzyme. This study, hypothesized and experimented on by several researchers, could have potentiality in the evolution of diabetes treatment if further carried out. By first testing out the phytochemicals with the best binding affinities in a laboratory using an assay kit, micro plate, and dosages of these phytochemicals. According to the results of the experimentation, the phytochemical with the most potential can be turned into a drug for the use of diabetes treatment. Patients with diabetes mellitus would take this medication, in the form of a pill, with their diet so that it could be put to work during the digestion process.
Keywords
Flexor hallucis longus tendon, Ruptured Achilles tendons, Lindholm technique, AOFAS
Introduction
The human Alpha Amylase enzyme is located our salivary glands as well as the pancreas. The alphaamylase enzyme works to speed up the reaction of converting starch from food into maltose units, which are then turned into glucose units by the enzyme “maltase”. The main difference between the amylase located in the pancreas and the salivary glands, is that pancreatic amylase works to digest more complex carbs (starches). Once food enters your salivated mouth, the salivary (amylase) enzyme, breaks down the starch into maltose (a two-unit glucose chain connected with a bond). The enzyme maltase then converts the maltose into glucose by breaking apart the two units. However, some complex carbs remain even after salivary amylase has started its starch conversion. Once the broken-up food travels down the esophagus and makes its way into the pancreas, the remaining starches get digested by the pancreatic amylase. Pancreatic fluid releases into the Enondrum, the first section in the small intestine. This is where the pancreatic a-amylase enzyme begins its starch digestion The a-amylase enzymes play a major role in the amount of blood glucose we have [1-3]. However, too much glucose in one’s blood stream is harmful and leads to a group of conditions known as Diabetes mellitus.
Diabetes mellitus refers to a group of conditions that causes high blood glucose (sugar) affecting over 10% of the U.S. population. The reason for such high rates, especially in this country, is due to excessive fast-food consumption, thus leading to obesity and high cholesterol levels. If not treated properly, the high blood sugar that is present in one's bloodstream could damage the blood vessels and the nerves present in the heart. Therefore, one with high blood sugar is twice as likely to get heart disease. As of now, there is no cure for diabetes, although it may be controlled. In most cases, insulin therapy is the most common treatment. Oral medication, especially for type 1 diabetes, isn't as common since it may inhibit the a-amylase enzyme while damaging other cells in the human body. For instance, the drug Jardiance has serious side effects that could potentially lead to other health conditions, making it less favorable and recommended for patients of diabetes [4-7].
Research has suggested that slowing down the reaction of starch- to- glucose conversion, one’s bloodstream would not have as much glucose accumulated. This can be done by preventing the catalysis reaction of the alpha-amylase enzyme. This is not an unfamiliar study, for several experiments have been performed in order to test which phytochemicals inhibit the a-amylase enzyme. The procedure of extracting plant extracts and using the compounds to test for a-amylase inhibition is well known and experimented. For instance, the Department of Pharmaceutical Sciences in Brazil has created a review about crude extracts and isolated compounds from plant sources to test for a- amylase inhibitory activity [8]. Although this study aims at discovering phytochemicals with structures that could potentially mask the site of the- amylase, an experimentation in a lab has not been done. Rather, molecular docking is the first step in reaching this objective.
Materials and Methods
Molecular docking allows us to test this protein- ligand binding. By downloading the structures of both an enzyme and a protein, anyone could perform molecular docking through software such as PyMOL [9], Autodock vina [10], and Open babel [11]. The point of this procedure is to see which ligands can potentially inhibit the protein being tested. Certain enzymes may catalyze chemical reactions that influence certain disorders in a negative way. Therefore, several oral medications that have been prescribed to patients with health complications, are made up of protein inhibitors.
In this study, molecular docking was performed by first downloading the human a-amylase enzyme structure from the Protein Data Bank (PDB) [12] and preparing the protein in Auto Dock by simplifying it so that there won’t be any variables serving as obstacles during the molecular docking. Afterwards, the file was then converted to the desired PDBQT format via Auto Dock Vina software. Subsequently, the active site of the protein was determined by highlighting the catalytic amino acid residues in AutoDock Vina and adjusting the sizes and coordinates of the grid box accordingly to fit the highlighted particles, after which a configuration document with the recorded grid coordinates was made. After the receptor was prepared, ligand determination was needed by researching for phytochemicals from plants that have already been medically proven to have effects on the alpha- amylase enzyme. These chemical compound structures were obtained from the National Center for Biotechnology Information (NCBI) [13] then converted to PDBQT format through the Open Babel Conversion Program. Once a configuration document was created for each ligand stating the grid information, ligand and receptor name, exhaustiveness, and energy range, then the ligands were docked by running the AutoDock Vina software to find the binding affinity between the active site of the protein and the ligand.
Once complete with the manual docking, high throughput was performed by supercomputers in the University of Texas at Austin Health Center (TACC drug discovery) [14]. Upon requesting for an account, the receptor can be uploaded, and the active site grid information may be inputted to start the screening process. TACC Drug Discovery runs millions of ligands and returns a document of the top 500 ligands that have the best (lowest) binding affinities.
After the molecular docking, a threshold was calculated by finding the standard deviation of the average binding affinities of already known alpha- amylase enzyme inhibitors. Then, the SMILES codes of the chemical compounds were inputted into the Mcule Toxicity Checker [15] to determine which phytochemicals have/ don’t have potentially toxic structures.
Results
Upon researching, 23 plants that have been proven to have effects on the alpha- amylase enzyme, were identified4. Out of these plants, 42 out of 109 phytochemicals that were ran, had binding affinities under the medically recommended threshold of -8 kcal/mol: Leucocyanidin, Quercitrin, Oleanolic Acid, Oxosteroid, Spinasterol, Lupeol, Barbaloin, Chrysophanol, Andrograpanin, Andrographolide, Deoxy Andrographolide, annonacin, anonaine, liriodenine, Isoquercitrin, Azadirachtin, Gedunin, Nimbin, Nimbiol, Nimbolide, Salannin, Verbascoside, Amentoflavone, Vitexin, chlorogenic Acid, Orientin, Naringin, Cucurbitacin, Isovitexin, Lutonarin, Saponarin, Genistin, Myricetin, Rutin, Theaflavin, Taxifolin, Cichoralexin, Lactucopicrin, Spicoside, Taraxacin, Taraxasterol, Hispidulin, and Linarin. Phytochemicals from the Dandelion (taraxesterol, Taraxacin, Chlorogenic Acid) had affinities that indicated the best binding with the proteins active site. Out of these chemicals, 7 are nontoxic: Linarin, Hispidulin, Amentoflavone, Isoquercitrin, Oleanolic Acid, Oxosteroid, and Cichoralexin. Oleanolic acid had the most potential out of these with a binding affinity of -11.72 kcal/mol. From the high throughput screening (TACC), over 10,000 Zinc’s had values under -8. Sixty- four chemicals have affinities below -7.01 kcal/ mol, the calculated threshold (Figures 1 to Figure 3).
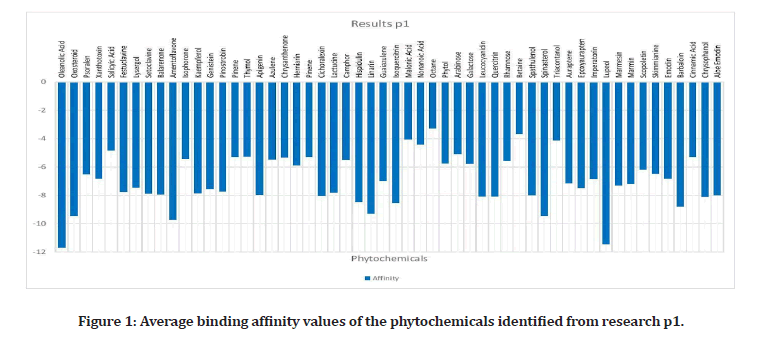
Figure 1: Average binding affinity values of the phytochemicals identified from research p1.
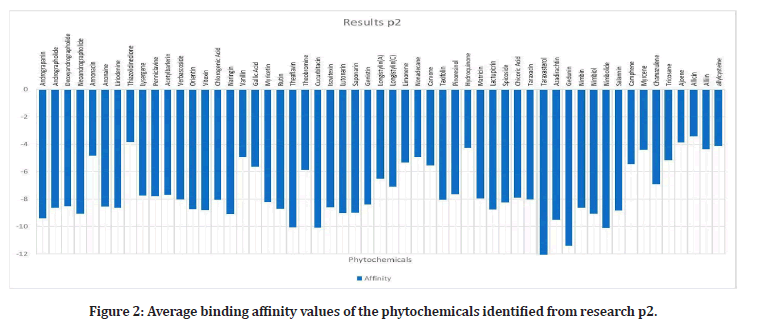
Figure 2: Average binding affinity values of the phytochemicals identified from research p2.
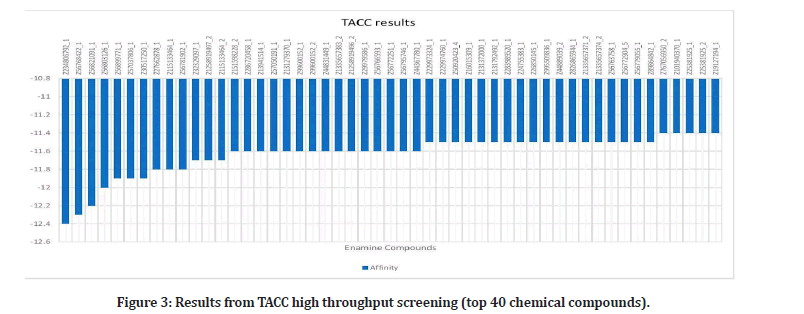
Figure 3: Results from TACC high throughput screening (top 40 chemical compounds).
Discussion
Diabetes Mellitus serves as one of the leading causes of death in the United States. Oral medication treatment with limited side effects, unfortunately, are uncommon. However, by preventing the catalysis reaction of starch digestion, blood glucose wouldn’t accumulate as quickly, thus decreasing blood sugar in patients with diabetes. This theory was carried out by researching and performing molecular docking.
Currently, on the market there is a diabetes treatment that serves as an inhibitor of the alpha-glucosidase enzyme, which also plays a role in the digestion of starch to glucose. This medication is composed of Acarbose [16]. Studies suggest that this inhibitor may have potentiality in inhibiting the alpha- amylase. As the alpha- amylase and alpha-glucoside enzyme do not have the same structure, the binding affinity between the active site of the alpha- amylase and Acarbose did not come out as desirable. By performing this research and molecular docking, previously unknown or unsure suggestions of certain phytochemicals as diabetes treatment were confirmed or contradicted.
The top phytochemicals, Taraxesterol and Oleanolic acid have potential binding affinities with the active site of the alpha- amylase. The phytochemical, Taraxesterol, is known to induce apoptosis. It is also of medical knowledge that it has strong affects against snake venom. In the long run, phytochemicals belonging to the plant Taraxacum, or dandelion, have the most promising binding affinity [17]. Oleanolic acid is a constituent of allium sativum, or garlic. Oleanolic acid is used in dermatology to prevent aging on the skin. Additionally, it is an anticancer phytochemical that is used for treatment of liver diseases [18].
This research revealed quite a handful of candidate drugs against the alpha amylase enzyme. Some of these drugs are toxic, meaning dosage and concentration of these compounds must be carefully monitored and calculated. Although requiring further experimentations for confirmation, the results from this study could serve as a guide for future references.
Conclusion
The phytochemicals ran through the docking software had impressive binding affinities with the active site of the alpha- amylase enzyme. In the health field, molecular docking is one of the first steps in developing an enzymeinhibitor drug that could treat a certain disease, or disorder. Therefore, these findings can be considered when establishing oral medication for diabetes mellitus treatment.
References
- Poovitha S, Parani M. In vitro and in vivo a-amylase and a-glucosidase inhibiting activities of the protein extracts from two varieties of bitter gourd (Momordica charantia L.). BMC Complement Altern Med 2016; 16:1-8.
- https://pdb101.rcsb.org/motm/74#:~:text=Alpha%2Damylase%20begins%20the%20process,two%20or%20three%20glucose%20units.
- Rudnitskaya A, Török B, Török M. Molecular docking of enzyme inhibitors. Biochem Mol Biol Educ 2010; 38:261-265.
- Kooti W, Farokhipour M, Asadzadeh Z, et al. The role of medicinal plants in the treatment of diabetes: A systematic review. Electron Phys 2016; 8:1832.
- https://mcule.com/apps/toxicity-checker/
- https://www.mayoclinic.org/diseases-conditions/diabetes/symptoms-causes/syc-20371444.
- Shah BS, Sartaj L, Ali F, et al. Plant extracts are the potential inhibitors of a-amylase: A review. Bioequivalence Bioavailab 2018; 5:270-273.
- https://www.jardiance.com/type-2-diabetes
- http://www.pymol.org/pymol
- Trott O, Olson AJ. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization and multithreading. J Comput Chem 2010; 31:455-461.
- O'Boyle NM, Banck M, James CA, et al. Open babel: An open chemical toolbox. J Cheminform 2011; 3:33.
- Berman HM, Westbrook J, Gilliland G, et al. The protein data bank. Nucleic Acids Res 2000; 28:235-242.
- Kim S, Chen J, Cheng T, Gindulyte A, et al. PubChem in 2021: New data content and improved web interfaces. Nucleic Acids Res 2021; 49:D1388-1395.
- https://drugdiscovery.tacc.utexas.edu/#/
- https://mcule.com/apps/toxicity-checker/?structure=MCULE-9778356861
- https://www.webmd.com/diabetes/guide/diabetes-general-treatments
- Wirngo FE, Lambert MN, Jeppesen PB. The physiological effects of dandelion (Taraxacum officinale) in type 2 diabetes. Rev Diabet Stud 2016; 13:113-131.
- Ayeleso TB, Matumba MG, Mukwevho E. Oleanolic acid and its derivatives: Biological activities and therapeutic potential in chronic diseases. Molecules 2017; 22:1915.
Google Scholar, Indexed at, Cross Ref
Google Scholar, Indexed at, Cross Ref
Google Scholar, Indexed at, Cross Ref
Google Scholar, Indexed at, Cross Ref
Google Scholar, Indexed at, Cross Ref
Google Scholar, Indexed at, Cross Ref
Google Scholar, Indexed at, Cross Ref
Google Scholar, Indexed at, Cross Ref
Google Scholar, Indexed at, Cross Ref
Author Info
Nursel Eski1, Cengiz Z Altuntas2, Harun Yilmaz3 and Senol Dane4*
1Student, Harmony School of Innovation, 22400 Grand Corner Dr Building C, Katy, TX 77494, USA2Texas Institute of Biotechnology Education & Research (TIBER), North American University, 11929 W Airport Blvd Houston, TX 77477, USA
3Education Department, North American University, 11929 W Airport Blvd Houston, TX 77477, USA
4Department of Physiology, Faculty of Basic Medical Sciences, College of Health Sciences, Nile University of Nigeria, Abuja, USA
Received: 07-Jul-2022, Manuscript No. jrmds-22-68824; , Pre QC No. jrmds-22-68824(PQ); Editor assigned: 09-Jul-2022, Pre QC No. jrmds-22-68824(PQ); Reviewed: 26-Jul-2022, QC No. jrmds-22-68824; Revised: 02-Aug-2022, Manuscript No. jrmds-22-68824 (R); Published: 09-Aug-2022
